Small molecule High Throughput Screen using AstraZeneca facilities
This is a really interesting opportunity. With the demise of the European Lead Factory access to a large, high quality screening deck is very limited.
The MRC now have a unique funding opportunity, a chance to screen against the AstraZeneca screening deck. https://www.ukri.org/opportunity/develop-new-approaches-to-small-molecule-medicine/?utmmedium=email&utmsource=govdelivery.
Funding priority in this round will be given to applications related to fibrosis or extracellular matrix targets.
Two funding opportunities a year with new thematic focus each round. Future areas will include:
- autoimmunity
- pain
- motor neuron disease
- mental health
- dementias (including Parkinson’s and Huntington’s)
- women’s health (including related to metabolic disorders)
Often the challenge for small groups is optimisation of the assay to make it suitable for HTS, so it is great this is also to be funded and AZ will provide technical advice
What will funded, costs related to the staff and consumable costs incurred at AstraZencea for the optimisation and execution of the High Throughput Screen (HTS). These include:
- £20,000 (100% FEC Exceptions) – Optimisation and establishment of an HTS
- £150,000 (100% FEC Exceptions) – Execution of the HTS
- cost of travel, accommodation and subsistence for a host institution researcher to work at AstraZeneca in Cambridge for three months (80% FEC)
- costs for elements of the screening cascade that cannot be undertaken at AstraZeneca and must be undertaken at the host organisation (80% FEC)
- minimal %FTE for the project lead (80% FEC)
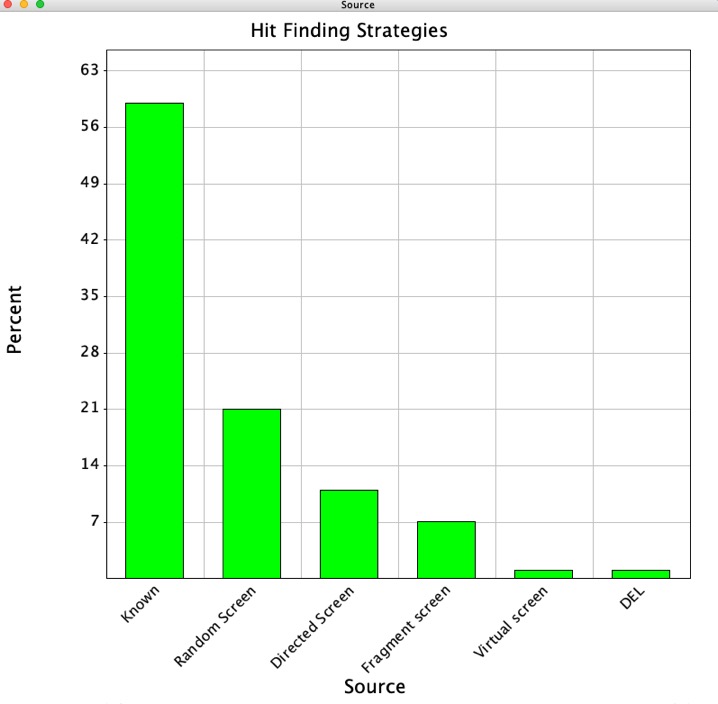
Updated Hit identification page
The Hit Identification page to include information from a great analysis published recently. An Analysis of Successful Hit-to-Clinical Candidate Pairs?" DOI.
Binding Sites are 3D
I've always found it interesting that whilst everyone recognises that protein binding sites are three dimensional (and chiral) there is a reluctance to have chiral centres in screening hits. This is despite examples were chiral centres aid affinity, selectivity and solubility. I suspect one of the concerns is the ease of follow up for any hits.
I've been working with Liverpool Chirochem to design a 3D rich, homochiral fragment screening library. The real beauty of this library is that the fragments can be easily expanded using validated chemistry in their parallel synthesis lab.
Once you have built a supply of these homochiral building blocks they can of course be put to many different additional uses, covalent fragments, DEL building blocks, and as building blocks for large virtual libraries. All of which can be supported by their parallel synthesis lab.
Submit a phenotypic assay
This sounds like an interesting opportunity.
Xcellomics is seeking novel, in vitro or ex vivo pathologically relevant cellular phenotypes that have the potential to be developed into small molecule High Content or CRISPR screens. We are encouraging members of the research community to submit proposals through the Xcellomics applications portal in the following disease areas….
Full details here https://www.xcellomics.com/calls
Xcellomics is a partnership between Exscientia and the University of Oxford Target Discovery Institute (TDI).
Ultra large Chemical Libraries
In a recent blog post Derek Lowe talked about "Virtual Screening Versus the Numbers" https://www.science.org/content/blog-post/virtual-screening-versus-numbers highlighting some of the issues around ultra large chemical libraries.
It seems quite timely that RSC CICAG is organising a meeting on Ultra Large Chemical libraries 10 August 2022 10:00-17:00, Burlington House, London, United Kingdom.
A decade ago a chemical library of a million compounds was considered large but over the last few years there has been a period of continuous growth in the size of both physical and virtual chemical libraries. As the libraries have grown the conventional search technologies have become unsustainable and new technologies are needed. This meeting will look at the challenges and solutions used to design, create, compare and search these ultra-large chemical libraries.
There are more details and registration here https://www.rsc.org/events/detail/73675/ultra-large-chemical-libraries.
It is now open for abstract submission (oral due by May 1st, posters June 2nd).
Registration fees
Delegate member early £95
Delegate non-member early £115
Delegate member std £120
Delegate non-member std £145
Student member early £65
Student non-member early £85
Student member std £90
Student non-member std £110
Find out more about the European Lead Factory
On Tuesday 30 November, members of the European Lead Factory (ELF) will participate in a webinar organised by the Young Scientists Network (YSN) of the European Federation of Medicinal Chemistry and Chemical Biology (EFMC).
The topic of the session is “Biological Testing of Hit Compounds”. Dr Vera Nies, Programme Manager at Lygature (the coordinating partner of the ELF) will give an introduction to the European Lead Factory. Her talk will be followed by a presentation entitled "Best practices in High Throughput Screening: ELF as an example", by Dr Steven van Helden of Pivot Park Screening Centre. The last ELF member presentation will be given by Dr Phil Jones of BioAscent, who will talk about the “Approaches Towards PAINSless Lead Generation”.
More details and registration here https://www.europeanleadfactory.eu/newsroom/elf-participate-efmc-ysn-webinar
AI4Proteins videos now online
On June 16/17 2021 RSC CICAG and AI3D held a joint meeting on Protein Structure Prediction. The full lineup of speakers, titles and abstracts can be found here.
Session 1: Session Chair: Professor Jeremy Frey (University of Southampton)
An AI solution to the protein folding problem: what is it, how did it happen, and some implications Professor John Moult (University of Maryland)
Session 2: Session Chair: Dr Melanie Vollmar (Diamond)
So you predicted a protein structure – What now? Dr Thomas Steinbrecher (Schrödinger)
Deep Learning enhanced prediction of protein structure and dynamics Dr Martina Audagnotto (AstraZeneca)
Fireflies-Lévy Flights algorithm for peptides conformational optimization Dr Zied Hosni (University of Sheffield)
Session 3: Session Chair: Dr Chris Swain (Cambridge MedChem Consulting)
How good are protein structure prediction methods at predicting folding pathways? Mr Carlos Outeiral Rubiera (University of Oxford)
Protein-Ligand Structure Prediction for GPCR Drug Design Dr Chris De Graaf (Sosei Heptares)
Session 4: Session Chair: Dr Márton Vass
Using icospherical input data in machine learning on the protein-binding problem Dr Ella Gale (University of Bristol)
Biological sequence design with machine learning Professor Debora Marks (Harvard University)
Session 5: Session Chair: Dr Simone Fulle (Novo Nordisk)
Lessons learned from generative models of biological sequences Professor Aleksej Zelezniak (Chalmers University of Technology)
DeepDock: a deep learning approach to predict ligand binding conformations Dr Oscar Méndez-Lucio (Janssen Pharmaceuticals)
Finding new in silico-based therapeutic strategies for IAHSP Dr Matteo Rossi Sebastiano (University of Turin)
Session 6: Session Chair: Professor Jonathan Goodman (University of Cambridge)
Designing molecular models by machine learning and experimental data Professor Cecilia Clementi (Freie Universität Berlin)
The “almost druggable” genome Professor Tudor Oprea (University of New Mexico)
Session 7: Session Chair: Dr Lucy Colwell (University of Cambridge)
General Effects of AI on Drug Discovery Dr Derek Lowe (Novartis)
Open Access Data: A Cornerstone for Artificial Intelligence Approaches to Protein Structure Prediction Professor Stephen Burley (RCSB PDB, Rutgers University, UCSD)
The videos of the presentations are now available on YouTube and you can access the playlist here https://www.youtube.com/playlist?list=PLBQwbn0mPhvWyTLnN6eFsbIwb5FByrs.
For those wanting a hype free insight into the impact AI might make on Drug Discovery then the presentation by Derek Lowe is well worth watching.
European Lead Factory call for proposals
The closing date for submissions to the European Lead Factory is Friday 28th May 2021.
Following the UK’s exit from the European Union on 31 January, we are pleased to confirm that the European Lead Factory still welcomes screening proposals from UK researchers.
This is a fantastic opportunity for researchers to get access to a top class high-throughput screening platform.
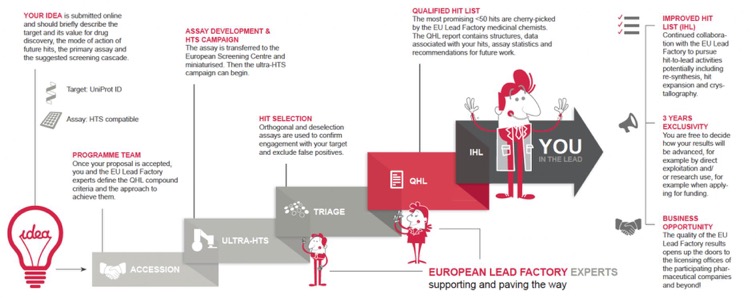
On this page, you can find the application forms, legal documents and all other relevant information required for your proposal submission.
The screening deck consists of around 535,000 compounds consisting of about 300,000 compounds from the original pharmaceutical companies, along with almost 190,000 compounds from custom design and synthesis. Furthermore, Grünenthal and Institut de Recherche Servier have provided around 50,000 new compounds not previously available to the European Lead Factory. Compounds from partners were selected for inclusion in the library based on several important quality criteria, as well as on the prerequisite that they are unique and not commercially available. Efforts were also taken to increase diversity, resulting in a top-quality compound library.
The single most important factor determining the likelihood of success of a project is the quality of the starting lead
Have you identified a disease mechanism that should be explored in a drug discovery project? Then you may have also considered developing a screening assay to find new chemical starting points that could serve as candidates for drug development.
In the next European Lead Factory webinar, taking place on Thursday 28 January, Dr Saman Honarnejad, Director of Drug Discovery at Pivot Park Screening Centre, will give a presentation on guidelines for HTS assay development, with an emphasis on methods to design, evaluate and improve conditions for biochemical and cellular assay formats.
The event will take place via Microsoft Teams on 28 January 2021 from 11:00-12:00 (CET). Discussions with the audience will form a key part of the webinar and time will be made available for questions and answers.
Registration is free using this link https://events.europeanleadfactory.eu/elf/elf-webinar/.
The European Lead Factory welcomes drug targets in all therapeutic areas. Interested? Submit your screening proposal now. The next deadline to apply will be in January 2021. Full details are on the ELF website.
I've been involved in a number of screening campaigns and I can confirm this is a high quality screening deck.
Still time to submit proposal to European Lead Factory
The closure date for the latest round of proposals for screening at the European Lead Factory is January 2021
The European Lead Factory welcomes drug targets in all therapeutic areas. Interested? Submit your screening proposal now. The next deadline to apply will be in January 2021.
Full details are on the ELF website.
I've been involved in a number of screening campaigns and I can confirm this is a high quality screening deck.
Proposals for European Lead Factory
The closure date for the latest round of proposals for screening at the European Lead Factory is January 2021
The European Lead Factory welcomes drug targets in all therapeutic areas. Interested? Submit your screening proposal now. The next deadline to apply will be in January 2021.
Also note
Following the UK’s exit from the European Union on 31 January, we are pleased to confirm that the European Lead Factory still welcomes screening proposals from UK researchers.
You can read more about the screening facility here https://www.europeanleadfactory.eu/node/353.
The details of how to submit are here https://www.europeanleadfactory.eu/how-submit/drug-target-assays/how-it-works.
I've written about the ELF here.

European Lead Factory Q&A
Interested in accessing a high quality high-throughput screening platform? Here is a chance to find out more about the European Lead Factory.
More details are here
The European Lead Factory (ELF) is a collaborative public-private partnership aiming to deliver novel lead molecules for drug discovery programs.
I've previously written about the ELF here.
Phenotypic Screening now offered by the European Lead Factory
The European Lead Factory has announced that it can now offer two types of phenotypic screening:
- A high-throughput, but “lower content” phenotypic approach that is suited to screening ELF’s entire compound collection, and
- A more complex “high content” screening approach using microscopy or flow cytometry to probe phenotype on a smaller subset of the compound collection
While low content assays can be live measurements or have fixed end points and involve well-averaged readouts, high content assays can be much more complex, based on live or fixed cells, multiple cell types and usually have more than one parameter as a readout. The complexity of the latter workflow makes it better suited to being performed on a smaller representative subset of the large collection.
Phenotypic screening historically has been the basis for the discovery of many drugs. Compounds are screened in cellular or animal disease models to identify compounds that cause a desirable change in phenotype. Only after the compounds have been discovered are efforts made to determine the biological targets of the compounds - a process known as target deconvolution.
Proposals for phenotypic screening approaches follow the normal review and selection process. A dedicated application form is available here.
The submission deadline for the next review and selection round is February 7, 2020.
European Lead Factory
Great news! European Lead Factory has restarted.
Pivot Park Screening Centre has successfully completed the first ultra-High Throughput Screening in IMI’s ESCulab project, and as such restarting the operations of the European Lead Factory. The screening on the European Compound Collection of ~500.000 compounds using a biochemical 1536-wells assay was finished within 4 days. Currently triaging of the UK owned program is ongoing within the Consortium, applying further biochemical and biophysical follow-up assays as well as the resynthesis of promising hits.
The programme is currently accepting proposals http://www.europeanleadfactory.eu/drug-target-assays.
European Lead Factory looking for novel screening programmes again
The European Lead Factory has been funded for another round of screening activities but under a new name European Screening Centre: unique library for attractive biology ESCulab. Over the next five years, the European Lead Factory will initiate 185 new drug discovery projects by screening medically relevant drug targets from European researchers, small and medium-sized enterprises and pharmaceutical industry against the ELF library of 550,000 unique chemical compounds.
The European Lead Factory was launched in 2013 and set up a joint collection of half a million compounds and a state-of-the-art high throughput screening centre. By the time the project ended last year, they had delivered results to researchers in universities, small biotechs and large companies across Europe, helping them to identify potential new drug candidates and breathing new life into a range of disease areas. In many cases, the seeds sown by the European Lead Factory resulted in new patents, partnering deals, and two start-ups. Now, a new IMI project, ESCulab will build on the work of the European Lead Factory. This means that researchers with drug targets can apply to screen the project’s compound collection for hits and get help developing any compounds further if they like. Jon de Vlieger, coordinator of the ESCulab consortium at Lygature, said: ‘It’s truly exciting to continue the onboarding of new and innovative proposals for screening and provide high quality starting points for drug discovery to academics and SMEs throughout Europe. In an effort to broaden our scope we are not only looking for target-based approaches, but now also enable phenotypic screens.’
You can apply here. If you don't have an assay in a format suitable for ultra high-throughput screening it is worth noting that the Wellcome Trust have small awards designed to help with the technology change required for HTS.
ChemBridge Macrocycle Library
Macrocycles offer a unique opportunity to address some of the more challenging drug targets and I've highlighted this on a couple of pages on the Drug Discovery Resources, here and here.
Macrocycles lie outside the usual "drug space" delineated by the Rule-of-5 and macrocycles can adopt different conformation in various media, hiding polar atoms or forming intramolecular hydrogen bonds, thus retaining good cell permeability and ADME properties
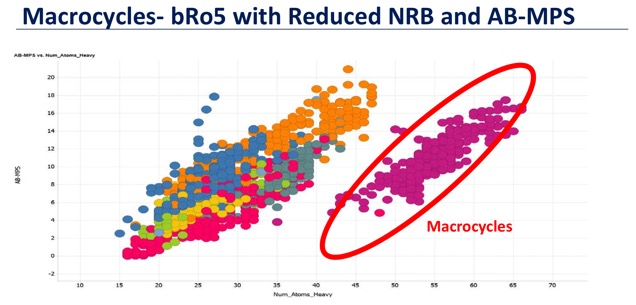
I recently got an email from ChemBridge highlighting a new 11,000 member Macrocyclic Library for screening. The general characteristics of compounds in the ChemBridge Macrocycle Library include:
- Molecular weight range up to 800
- Primary ring size ranging from 11 to 27 atoms
- Heterocyclic primary rings
- Scaffolds with and without peptidic backbone elements as part of the macrocyclic ring
- Scaffolds with and without fused rings as part of the primary macrocyclic ring
Developing an assay for high-throughput screening
SULSA’s Assay Development Fund is actively recruiting innovative molecular targets for which there is a strong rationale for therapeutic potential.
Development of a high-throughput assay provide access to the various drug discovery initiatives that are available to the academic community, i.e. MRC DPFS, Bayer G4Targets, Wellcome Trust translational fund, the European Lead Factory, AZ innovation portal etc.
How many compounds do you select from virtual screening?
Whilst high-throughput screening (HTS) has been the starting point for many successful drug discovery programs the cost of screening, the accessibility of a large diverse sample collection, or throughput of the primary assay may preclude HTS as a starting point and identification of a smaller selection of compounds with a higher probability of being a hit may be desired. Directed or Virtual screening is a computational technique used in drug discovery research designed to identify potential hits for evaluation in primary assays. It involves the rapid in silico assessment of large libraries of chemical structures in order to identify those structures that most likely to be active against a drug target. The key question is then how many molecules do you select from your virtual screen?
Whilst virtual screening is certainly less expensive than high-throughput screening it is not free, even an in house academic cluster has an overhead (probably equating to > $10,000 per virtual screen). So with that investment how much would you invest in actual compounds?
Trust but verify
There is an article in Nature describing a collection of problems that have arisen from incorrect chemical structures in biological screens DOI. I'm slightly surprised that this is regarded as newsworthy, but I guess it serves as a timely reminder.
The data from screening campaigns invariably contains errors
- It is often a single point assay
- Quality and diversity of Sample Collection is variable
- Compounds may interfere with the detection system
- False positives due to aggregation
- High density plates can result in cross contamination, edge effects
There is a very simple mantra you should adopt when analysing screening data "Trust but verify".
- Check compounds that were found active against the selected target are re-tested using the same assay conditions used during the HTS.
- Does a resynthesised (not repurchased) show the same activity
- Dose response curve generation: an IC50 or EC50 value is then generated, does it have a reasonable slope? Uneffected by incubation time.
- Are related analogues available, check for genuine Structure-Activity Relationships

There is a strategy for the analysis of HTS data in the Drug Discovery Resources.
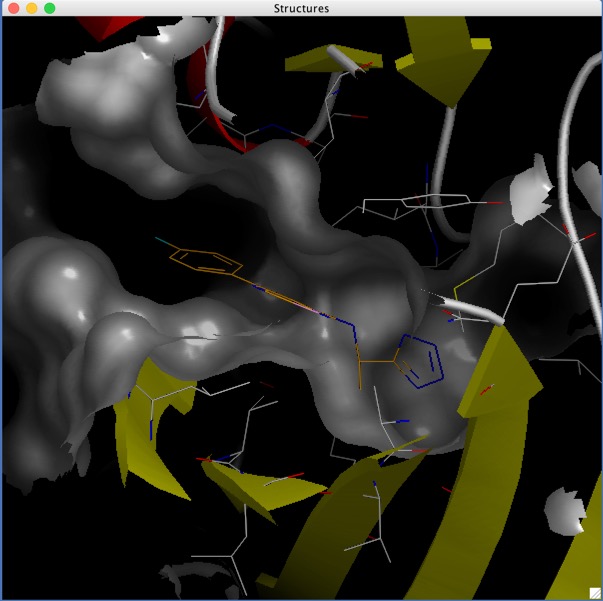
Virtual Screening Pages Updated
I've updated the pages describing virtual screening, in particular the docking section.
Heart on a chip
Cardiotoxicity is a major hurdle for all drug discovery programs and so I'm always interested in ways that any potential liabilities can be flagged without the need to go into whole animals. A recent publication highlights progress in developing a novel model system.
Human induced pluripotent stem cell-derived cardiomyocytes were cultured as a model system, and used to validate the platform with an excitation–contraction decoupling chemical. Preliminary data using the platform to investigate the effect of the drug norepinephrine are combined with computational efforts. This platform provides a quantitative and predictive assay system that can potentially be used for comprehensive assessment of cardiac toxicity earlier in the drug discovery process. DOI.
The European Lead Factory Works!
The European Lead Factory (ELF) is a collaborative public-private partnership aiming to deliver novel lead molecules for drug discovery programs. When the consortium was formed around 5 years ago there was a lot of scepticism about whether a group of 30 partners rating from large Pharma companies to small academic groups could ever agree on a legal framework that would allow the ELF to function. In a addition, in an industry where confidentiality was critical to maintaining intellectual property the idea that a group of large Pharma companies would share their sample collections often regarded as the "Crown Jewels" seemed impossible. However I was at the European Lead factory Stakeholder Meeting (24-25 April 2017) and it is clear that is has been a success.
European Lead factory Newsletter
The latest issue of the European Lead Factory Newsletter has just been published.
Highlights include the European Lead Factory recently reached the milestone of having received over 1,000 chemical library proposals for consideration. To date, 143,529 novel compounds, out of the 200,000 prospected compounds for the Public Compound Collection, have been synthesized, which means that the ELF compound collection grows daily by approximately 250 novel compounds to eventually constitute the 500,000 Joint European Compound Library (JECL).
Parkinson’s UK and the University of Sheffield have launched a joint venture biotech company, Keapstone Therapeutics. Parkinson’s UK has allocated 1 million GBP over the next sixteen months to further develop compounds that boost the internal cellular defence mechanisms against oxidative stress. These compounds were discovered by European Lead Factory.
The Ecstasy and Agony of Assay Interference Compounds
There is an editorial in ACS Central Science DOI that I would encourage everyone involved in hit identification to read.
A couple of quotes will give you an idea of the content
Alarmingly, up to 80–100% of initial hits from screening can be artefacts if appropriate control experiments are not employed.
it is important to realize that no PAINS-containing drug has ever been developed starting from a protein-reactive PAINS target-based screening hit
They also emphasise the critical need for experimental validation for any screening hit.
Such validation experiments include classic dose response curves, lack of incubation effects, imperviousness to mild reductants, and specificity versus counter-screening targets. If a molecule is flagged as a potential PAINS or aggregator using published patterns but is well-behaved by these criteria, it may be a true, well-behaved ligand. Ultimately, genuine SAR combined with careful mechanistic study provides the most convincing evidence for a specific interaction. Covalent and spectroscopic interference molecules act via specific physical mechanisms, for which controls are known. Colloidal aggregation, fortunately, is readily identified by rapid mechanistic tests and by counter-screening.
In addition you need to consider compound identify and purity, reproducing the activity with an authentic sample is essential.
Whilst time-consuming this validation work will save a fortune in the future.
Peptidyl-Prolyl cis-trans Isomerase (PPIase) assays
Having worked wit Selcia on a number of projects I always keep an eye out for news on their work on Peptidyl-Prolyl cis-trans Isomerases (PPIase). These are very interesting class of enzymes whose principal function is to catalyse the cis-trans isomerisation of the X-Pro peptide bonds in polypeptide chains (where X is any amino acid). This transformation is thought to be a mechanism to modulate protein function.
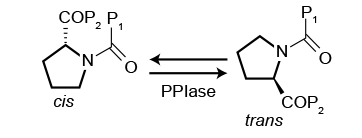
PPIase enzyme targets are of increasing interest in drug discovery due to the extensive potential of small molecule inhibitors in a range of therapeutic areas, including infection, inflammation, cancer and neuroprotection.
Selcia have now expanded the range of Peptidyl-Prolyl cis-trans Isomerase (PPIase) assays they can offer.
The SULSA Assay Development Fund:
I've worked with a couple of academic groups who have a very interesting target but no small molecule leads. Whilst there are several places offering high-throughput screening the stumbling block is often development of a robust screening assay running in a high density format. Most academic groups have little experience in developing such an assay and to be honest will need to do so very infrequently.
The SULSA Assay Development Fund: accelerating translation of new biology from academia to pharma DOI describes efforts to support the development of such assays, work carried out in collaboration with the European Lead Factory in Newhouse.
However, many scientifically interesting, novel molecular targets lack associated high-quality, robust assays suitable for hit finding and development. To bridge this gap, the Scottish Universities Life Sciences Alliance (SULSA) established a fund to develop assays to meet quality criteria such as those of the European Lead Factory. A diverse project portfolio was quickly assembled, and a review of the learnings and successful outcomes showed this fund as a new highly cost-effective model for leveraging significant follow-on resources, training early-career scientists and establishing a culture of translational drug discovery in the academic community.
European Lead Factory technical acceptance criteria
- Minimally 384 wells (max 30 μl)
- Homogenous assay (no washing steps)
- Defined endpoint
- Z-prime >0.6
- Readout stability >1 h
- S/B signal >3
- Incubation times <4 h
- DMSO tolerance: minimum 0.5%
- All reagents minimally 8 h stable
- Recombinant proteins(s) >80% pure
Perhaps this quote sums things up very nicely.
‘We only got so far in optimising our assay because in my opinion we do not have either the equipment or the personnel with the skill set required to do this efficiently. So being able to come and work at Newhouse where you are then surrounded by people who can identify quickly where assay improvements should be made and then pass on the knowledge of how to set about applying these changes (as well as gaining experience with various pieces of equipment) was invaluable.’
More details are available here http://www.sulsa.ac.uk/sites/sbsweb2.bio.ed.ac.uk.sulsa/files/downloads/SULSAassaydevelopmentfundguidancenotes4.pdf
The Wellcome Trust also provides Seed/Pathfinder awards that could cover similar projects.
Another European Lead Factory success
We are now starting to see some of the results of the screening of academic projects at the European Lead factory that was initiated in 2013.
Dr Mahlapuu’s group, based at the University of Gothenburg, first identified a new target which could be used to reverse metabolic complications in type 2 diabetes. With the help of the European Lead Factory experts, she then screened the Joint European Compound Library of the then 320,000 industry compounds and identified a set of selective and potent small molecules which interfere with this target.
They have now formed a spinout company to develop these leads. ScandiCure has received 1 MSEK from the 2016 SWElife program to continue the development of first-in-class anti-diabetic drug based on small molecule antagonists of a novel key mediator - serine/threonine protein kinase 25 (STK25).
ELF and antibiotic resistance programme
The latest news report from the European Lead Factory highlights work targeting antimicrobial resistance in collaboration with Professor Chris Schofield (University of Oxford). The high throughput screen of >300,000 compounds and initial triaging provided 50 qualified hits.
Multiple series of compounds were validated through resynthesis, biochemical and biophysical profiling at the European Screening Centre site in Newhouse, complemented with ligand–protein crystallography and antimicrobial evaluation at University of Oxford
Great to see projects like this move forward, demonstrates how important the ELF is in providing hits for academic groups/small companies.
European Lead Factory Update
The latest newsletter from the European Lead Factory has just been published and highlights a number of notable achievements.
As a result of the screening campaigns >3000 qualified hits have been awarded to private and public target owners. 72 public target programmes have been accepted, 48 high throughput screens finished and 41 hit lists with associated data reports handed over to the target owners. >150 bespoke assays have been developed in order to extract the most interesting hits for public programmes.
There are now 450,000 compounds in the compound library of which 120,000 are novel compound specifically synthesised for the ELF.
You can read more details here https://www.europeanleadfactory.eu/results/
The European Lead Factory is a collaborative public-private partnership aiming to deliver innovative drug discovery starting points. Having established the first European Compound Library and the first European Screening Centre, the EU Lead Factory aims to give free access to up to 500,000 novel compounds, a unique industry-standard uHTS platform, and much more.
We are now starting to see publications describing these endeavours..
https://www.europeanleadfactory.eu/results/publications/scientific-articles/
CO-ADD web portal
The CO-ADD web portal is now live You can now submit your free antimicrobial screening request, download all forms and access your primary screening, cytotoxicity, hit confirmation and hit validation reports online on the new secure CO-ADD user portal.
CO-ADD (Community for Open Antimicrobial Drug Discovery) is a not-for-profit initiative led by academics at The University of Queensland. Our goal is to screen compounds for antimicrobial activity for academic research groups for free. We aim to help researchers worldwide to find new, diverse compounds to combat drug-resistant infections.
Community for Open Antimicrobial Drug Discovery
A little while ago I mentioned The Community for Open Antimicrobial Drug Discovery effort to provide free compound screening against a variety of infective agents. .
Primary Screening of a 1mg sample Test against key ESKAPE pathogens, E. coli, K. pneumoniae, A. baumannii, P. aeruginosa, S. aureus (MRSA), as well as the fungi C. neoformans and C. albicans, at a single concentration.
Hit Confirmation:- Confirm activity with minimum inhibitory concentration and counterscreen for cytotoxicity and membrane interaction.
Hit Validation:- Test the positive hit against a broader panel of microbes and evaluate the basic drug qualities of actives. CO-ADD will screen your compounds for free and make no claim to IP. The linked flyer gives full details
You can also read more details in this Nature article DOI
It looks like they have achieved an important milestone!
Thanks to our research community we have received 100,000 compounds from 30 countries in 15 months! Make a difference: clear the fridge, empty the shelves and send through your compounds for free antimicrobial screening against 5 bacteria and 2 fungi. We would also like to acknowledge the contribution of the French National Chemical Library that has safely arrived in Brisbane last week!
So if you have compounds sitting in the back of cupboards why not send them to be tested
Drug Discovery Resources Update
I've updated the hit identification section of the Drug Discovery Resources. In particular I've added to the high-throughput screening analysis including more information on PAINS (Pan Assay Interference Compounds) first described by Baell et al DOI and subsequently summarised in an excellent Nature comment.
Academic researchers, drawn into drug discovery without appropriate guidance, are doing muddled science. When biologists identify a protein that contributes to disease, they hunt for chemical compounds that bind to the protein and affect its activity. A typical assay screens many thousands of chemicals. ‘Hits’ become tools for studying the disease, as well as starting points in the hunt for treatments.
These molecules — pan-assay interference compounds, or PAINS — have defined structures, covering several classes of compound. But biologists and inexperienced chemists rarely recognize them. Instead, such compounds are reported as having promising activity against a wide variety of proteins. Time and research money are consequently wasted in attempts to optimize the activity of these compounds. Chemists make multiple analogues of apparent hits hoping to improve the ‘fit’ between protein and compound. Meanwhile, true hits with real potential are neglected.
Also added a page on Aggregators. Promiscuous inhibition caused by small molecule aggregation is a major source of false positive results in high-throughput screening. To mitigate this, use of a nonionic detergent such as Triton X-100 or Tween-80 has been studied, which can disrupt aggregates, and is now common in screening campaigns DOI.
An Aggregation Advisor for Ligand Discovery
Aggregation is a regular concern when evaluating potential hits from screening and a recent paper "An Aggregation Advisor for Ligand Discovery" DOI attempts to provide an insight into this phenomenon, in addition they provide a useful web-based tool http://advisor.bkslab.org that provides a free service to advise whether molecules may aggregate under biological assay conditions.
Antibiotic Screening
A little while ago I mentioned The Community for Open Antimicrobial Drug Discovery effort to provide free compound screening against a variety of infective agents. I now have a few more details of what you might be able to access for a 1mg sample.
Primary Screening:-
Test against key ESKAPE pathogens, E. coli,
K. pneumoniae, A. baumannii, P. aeruginosa,
S. aureus (MRSA), as well as the fungi C. neoformans and C. albicans, at a single concentration.
Hit Confirmation:-
Confirm activity with minimum inhibitory concentration and counterscreen for cytotoxicity and membrane interaction.
Hit Validation:-
Test the positive hit against a broader panel of microbes and evaluate the basic drug qualities of actives.
CO-ADD will screen your compounds for free and make no claim to IP
European Lead Factory
p>The European Lead Factory has just announced that an additional 50,000 new compounds have been added to their screening collection. This brings the collection up to 350,000 compounds and sets them well on the way to their 500,000 target.
I've been involved in a couple of projects that have made use of this high-throughput screening facility and I've been impressed with the quality and diversity of the hits generated.
The European Lead Factory was established to promote the discovery of novel lead compounds, suitable for subsequent optimization either to drug candidates or to high‐quality pharmacological tools for the experimental validation of targets.
If you have a target you want to screen against you can submit a proposal online. For an academic or small company this is an interesting way to identify novel starting points for a medicinal chemistry program.
Protein-Protein Interactions
I’ve just added a section on Protein-Protein interactions.
These seem to be an increasingly popular target class and I’ hope to expand on it as we find out more.
And now for something completely different
In the last few updates I’ve concentrated on Fragment-Based-Screening, where by screening a small set of low molecular weight fragments it is possible to obtain hits that can then be expanded. The latest update highlights the use of encoded libraries, here vast libraries (>>1M compounds) can be screened the hits isolated and identified by virtue of a DNA tag.
From the early “proof-of-principle” peptide libraries this area has now extended into a variety of interesting chemistries.
European Lead Factory
I’m delighted to see the announcement about the European Lead Factory, hopefully this will be a huge asset for Drug Discovery. http://www.nature.com/news/europe-bets-on-drug-discovery-1.12372
Two sites shuttered by the pharmaceutical giant Merck, one in Scotland and one in the Netherlands, will soon be humming again with the work of drug discovery. But the hum will not be business as usual. It will be the sound of a public–private consortium placing a high-stakes wager: a nearly €200-million (US$271-million) bet that it can boost a languishing pharmaceutical sector by fusing academic innovation with industrial-scale screening, using robots to test chemicals for biological activity….Any academic group or company can also propose assays to test molecules in the library for biological activity. Lead-factory scientists will run these assays free of charge and confirm any promising results, working mainly in laboratory space closed by Merck in 2011 at Oss in the Netherlands. Follow-up work will be done at the University of Dundee in Scotland. Results will be provided confidentially to the groups that proposed the assays so that they can pursue further work and publications.
Building a Screening Collection
I’ve just updated the page on building a screening collection and added links to recent useful publications.
Broad Coverage of Commercially Available Lead-like Screening Space with Fewer than 350,000 Compounds, Jonathan Baell DOI
Metal Impurities Cause False Positives in High-Throughput Screening Campaigns, Johannes C. Hermann et al DOI
Malaria HTS results
Recently two groups have reported the results of screening campaigns, GlaxoSmithKline (GSK) published ( doi:10.1038/nature09107 ) the results of screening nearly 2 million compounds from their chemical library for inhibitors of P. falciparum, of which 13,533 were confirmed to inhibit parasite growth by at least 80% at 2 uM. The chemical structures and associated data were made public to encourage additional drug lead identification efforts and further research into this disease. In a second paper ( doi:10.1038/nature09099 ) a library containing 309,474 unique compounds, “designed at the scaffold level to provide diverse, comprehensive coverage of bioactive space”, was screened against Plasmodium falciparum strain 3D7 at a fixed concentration of 7 uM and afforded 1536 hits.
I thought I'd contribute to this generous effort by calculating a number of descriptors and properties for the molecules and then cluster the molecules in a variety of ways to help analysis and provide the data for download. You can read about it here.
