More COVID-19 MPro Activity Data
One of the best drug targets among coronaviruses is the main protease (Mpro), this enzyme is essential for processing the polyproteins that are translated from the viral RNA and the recognition sequence at most sites is Leu-Gln↓(Ser,Ala,Gly) and since no human enzymes have similar specificity inhibitors should be very specific. Mpro is a papain-like protease cysteine protease.
I've previously described the fragment hits from a fragment screen against crystals of the main protease (MPro) of SARS-CoV-2, the virus that causes COVID-19. Full details of the screening effort are described here https://www.diamond.ac.uk/covid-19/for-scientists/Main-protease-structure-and-XChem/Downloads.html
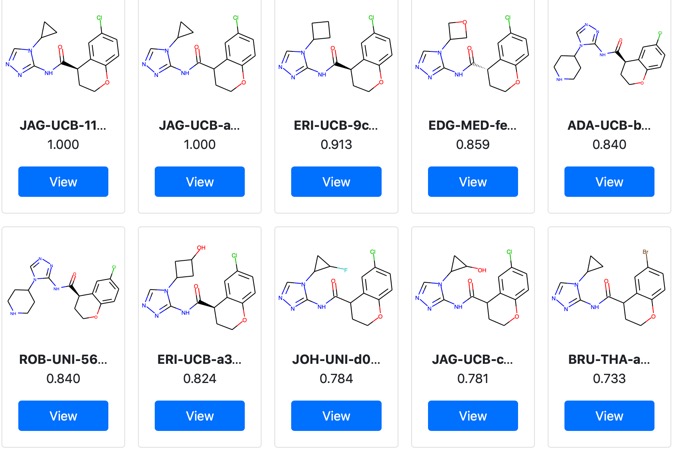
Additional biological results from project moonshot are now available. You can browse the data here https://postera.ai/covid/activity_data.
These results contain a significant milestone with the identification of the first sub micromolar non-covalent inhibitor.
JAG-UCB-a3ef7265-20 has been titrated twice now and has an IC50 of 0.6 uM.
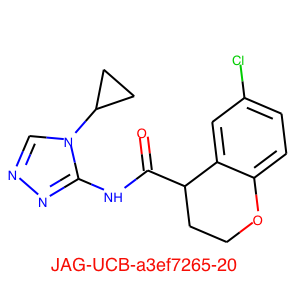
This compound is a racemic mixture and the synthesis of the individual enantiomers is underway, if the activity predominantly lies with a single enantiomer we could see a further improvement in activity. The original submission was based on a pharmacophore search of Enamine based on amino-pyridine hits. I highlight this to underline the importance of simple descriptor-based searches, they are often highly competitive with sophisticated docking studies and require orders of magnitude less compute resources.
Since this research is being conducted in the public domain a number of other people have been able to contribute further ideas based on this exciting discovery.