EMBL-EBI User Survey 2024
I suspect many of the readers of this site have used some of EMBL-EBI resources and tools are some time.
EMBL’s European Bioinformatics Institute maintains the world’s most comprehensive range of freely available and up-to-date molecular data resources.
https://www.ebi.ac.uk/services/data-resources-and-tools.
These of course include the Alphafold database. ChEMBL, PDB in Europe and much. Much more.
In order to support continued funding they do regular user surveys, and the latest is now online.
https://www.surveymonkey.com/r/HJKYKTT
It takes around 15 mins to complete so please take a tea break and fill it in.
AlphaFold Protein Structure Database in 2024
A recent publication describes the continued evolution of the AlphaFold Protein Structure Database created by EMBL-EBI and DeepMind. From an initial 300K structures it now contains 214 million predicted protein structures.
You can read the paper here DOI.
The AlphaFold Database Protein Structure Database (AlphaFold DB, https://alphafold.ebi.ac.uk) has significantly impacted structural biology by amassing over 214 million predicted protein structures, expanding from the initial 300k structures released in 2021. Enabled by the groundbreaking AlphaFold2 artificial intelligence (AI) system, the predictions archived in AlphaFold DB have been integrated into primary data resources such as PDB, UniProt, Ensembl, InterPro and MobiDB. Our manuscript details subsequent enhancements in data archiving, covering successive releases encompassing model organisms, global health proteomes, Swiss-Prot integration, and a host of curated protein datasets. We detail the data access mechanisms of AlphaFold DB, from direct file access via FTP to advanced queries using Google Cloud Public Datasets and the programmatic access endpoints of the database. We also discuss the improvements and services added since its initial release, including enhancements to the Predicted Aligned Error viewer, customisation options for the 3D viewer, and improvements in the search engine of AlphaFold DB.
Major user experience update in AlphaFold Database
Just saw this.
The AlphaFold Protein Structure Database, a result of a collaborative effort between Google DeepMind and EMBL’s European Bioinformatics Institute (EMBL-EBI), has released an exciting update to its web pages, providing users with an enhanced experience. This update marks a significant step in facilitating the use of AlphaFold structure data.
One of the most interesting updates are the improvements to the 3D viewer Mol*.
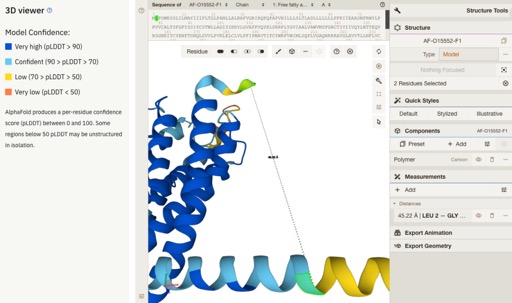
Full details are here https://www.ebi.ac.uk/about/news/updates-from-data-resources/alphafold-database-ux-update/.